A new paper by Emilia Huerta-Sánchez and colleagues was published in Nature yesterday: Altitude adaptation in Tibetans caused by introgression of Denisovan-like DNA.
Abstract
As modern humans migrated out of Africa, they encountered many new environmental conditions, including greater temperature extremes, different pathogens and higher altitudes. These diverse environments are likely to have acted as agents of natural selection and to have led to local adaptations. One of the most celebrated examples in humans is the adaptation of Tibetans to the hypoxic environment of the high-altitude Tibetan plateau. A hypoxia pathway gene, EPAS1, was previously identified as having the most extreme signature of positive selection in Tibetans, and was shown to be associated with differences in haemoglobin concentration at high altitude. Re-sequencing the region around EPAS1 in 40 Tibetan and 40 Han individuals, we find that this gene has a highly unusual haplotype structure that can only be convincingly explained by introgression of DNA from Denisovan or Denisovan-related individuals into humans. Scanning a larger set of worldwide populations, we find that the selected haplotype is only found in Denisovans and in Tibetans, and at very low frequency among Han Chinese. Furthermore, the length of the haplotype, and the fact that it is not found in any other populations, makes it unlikely that the haplotype sharing between Tibetans and Denisovans was caused by incomplete ancestral lineage sorting rather than introgression. Our findings illustrate that admixture with other hominin species has provided genetic variation that helped humans to adapt to new environments.
Xin Yi et al. (2010) were the ones who identified the EPAS mutation. They estimated that the Tibetan and Han Chinese populations diverged from one another as recently as 2750 years ago, and that the EPAS variant modifying the haemoglobin concentration underwent a very fast positive selection in Tibetans after that. In other words, the Denisovan version of the EPAS gene could have been passed to modern humans over 30,000 years ago (before Denisovans went extinct), but its frequency only soared after some East Asians settled in Tibet. This means that this Denisovan gene should also be found at lower frequency among all East Asian populations, and by extension perhaps also among Native Americans, Central Asians, Siberians, and perhaps even at trace frequencies among Europeans and Middle Easterners.
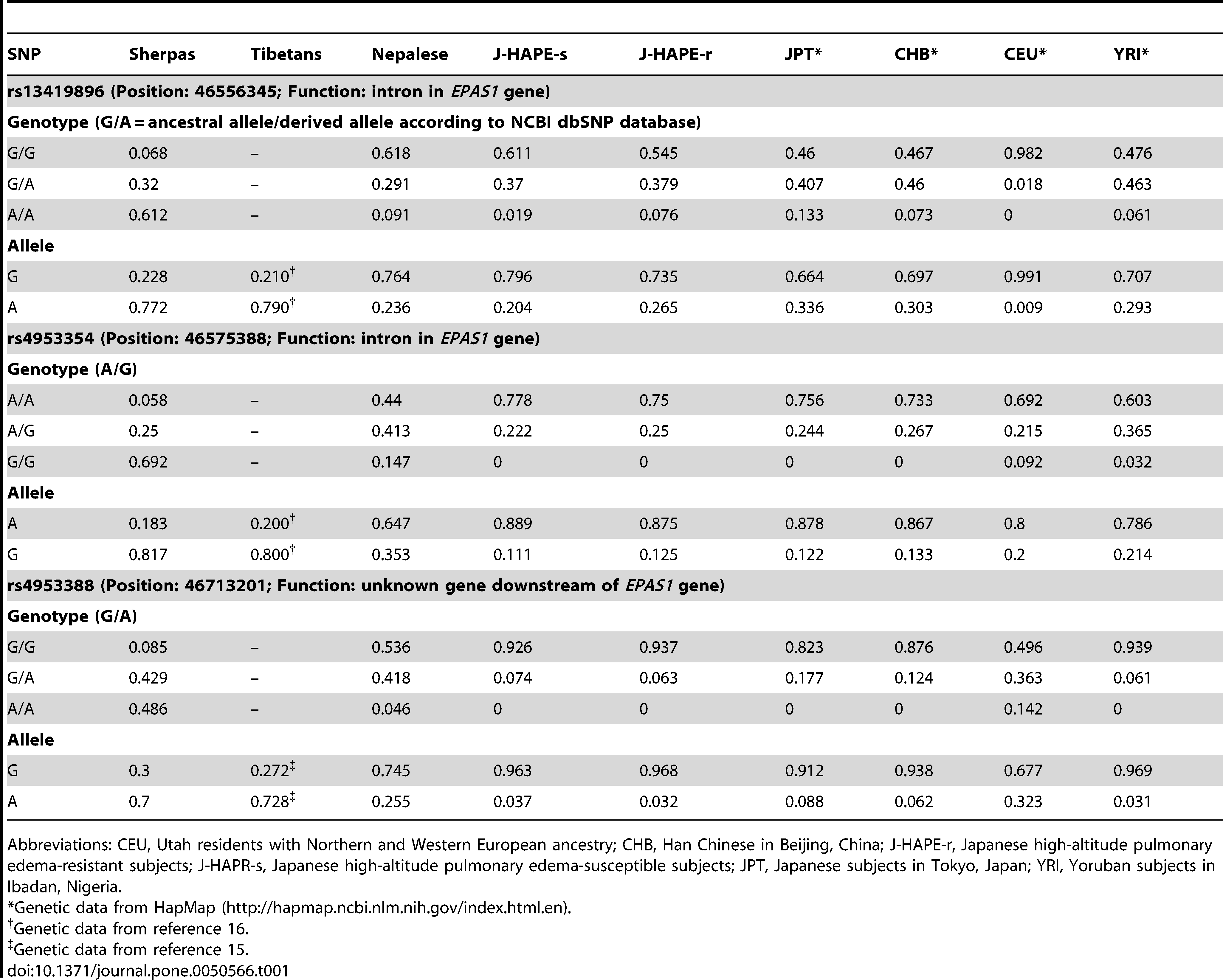
Hanaoka et al. (2012) confirmed the association of EPAS1 with adaptation to high-altitude hypoxia in the Sherpa population. They mention three significant SNPs (rs13419896, rs4953354, and rs4953388) in the EPAS1 gene and compare frequencies in the Sherpa, Tibetan, Nepalese, Han Chinese, Japanese, European and African populations. It confirms that the derived alleles are found in all populations.
Oddly enough only the first SNP is more frequent in East Asians. In fact the Japanese and Han Chinese completely lack the homozygous derived allele for the 2nd and 3rd SNPs. If these alleles all came from Denisovans, how did they end up being more frequent in Europeans and Africans than in East Asians (except Himalayan populations) ?
I wonder if the EPAS1 variant for increased haemoglobin concentration could also have been positively selected in other mountainous populations (Caucasus, Alps, Pyrenees) or in very hot climates where the air is also thinner.
Abstract
As modern humans migrated out of Africa, they encountered many new environmental conditions, including greater temperature extremes, different pathogens and higher altitudes. These diverse environments are likely to have acted as agents of natural selection and to have led to local adaptations. One of the most celebrated examples in humans is the adaptation of Tibetans to the hypoxic environment of the high-altitude Tibetan plateau. A hypoxia pathway gene, EPAS1, was previously identified as having the most extreme signature of positive selection in Tibetans, and was shown to be associated with differences in haemoglobin concentration at high altitude. Re-sequencing the region around EPAS1 in 40 Tibetan and 40 Han individuals, we find that this gene has a highly unusual haplotype structure that can only be convincingly explained by introgression of DNA from Denisovan or Denisovan-related individuals into humans. Scanning a larger set of worldwide populations, we find that the selected haplotype is only found in Denisovans and in Tibetans, and at very low frequency among Han Chinese. Furthermore, the length of the haplotype, and the fact that it is not found in any other populations, makes it unlikely that the haplotype sharing between Tibetans and Denisovans was caused by incomplete ancestral lineage sorting rather than introgression. Our findings illustrate that admixture with other hominin species has provided genetic variation that helped humans to adapt to new environments.
Xin Yi et al. (2010) were the ones who identified the EPAS mutation. They estimated that the Tibetan and Han Chinese populations diverged from one another as recently as 2750 years ago, and that the EPAS variant modifying the haemoglobin concentration underwent a very fast positive selection in Tibetans after that. In other words, the Denisovan version of the EPAS gene could have been passed to modern humans over 30,000 years ago (before Denisovans went extinct), but its frequency only soared after some East Asians settled in Tibet. This means that this Denisovan gene should also be found at lower frequency among all East Asian populations, and by extension perhaps also among Native Americans, Central Asians, Siberians, and perhaps even at trace frequencies among Europeans and Middle Easterners.
Hanaoka et al. (2012) confirmed the association of EPAS1 with adaptation to high-altitude hypoxia in the Sherpa population. They mention three significant SNPs (rs13419896, rs4953354, and rs4953388) in the EPAS1 gene and compare frequencies in the Sherpa, Tibetan, Nepalese, Han Chinese, Japanese, European and African populations. It confirms that the derived alleles are found in all populations.
Oddly enough only the first SNP is more frequent in East Asians. In fact the Japanese and Han Chinese completely lack the homozygous derived allele for the 2nd and 3rd SNPs. If these alleles all came from Denisovans, how did they end up being more frequent in Europeans and Africans than in East Asians (except Himalayan populations) ?
I wonder if the EPAS1 variant for increased haemoglobin concentration could also have been positively selected in other mountainous populations (Caucasus, Alps, Pyrenees) or in very hot climates where the air is also thinner.
